We operate one of the largest melanoma and immunotherapy cohorts in the world — combining clinical records, digital pathology, genomics, and spatial imaging to build AI models that directly improve cancer care.
Most melanoma deaths occur in patients initially diagnosed with early-stage (Stage I–II) disease who later experience recurrence. Our machine learning platform — trained on 1,720 melanomas across MGB and DFCI — applies AI in skin disease to predict which patients face the highest recurrence risk, exemplifying dermatology precision medicine in clinical informatics.
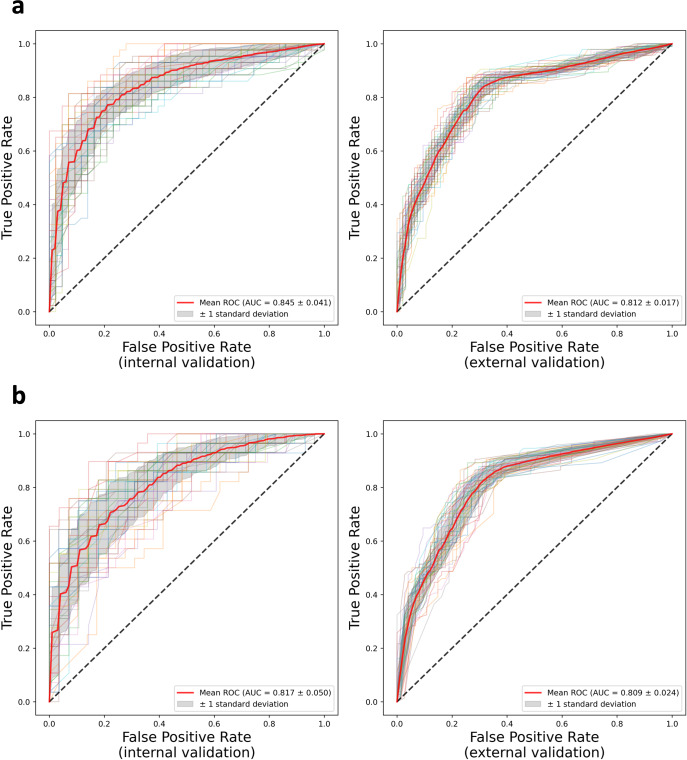
Our ML models integrate digital pathology, imaging, and clinical data using advanced computer vision techniques, achieving AUROC 0.845 internally and 0.812 externally — outperforming AJCC staging in predicting recurrence of non-metastatic melanoma. These models are currently being scaled across multiple external institutions as part of our clinical informatics in melanoma program.
Current TNM staging leaves oncologists with limited tools to identify which early-stage patients will relapse — our models provide individualized recurrence risk at the time of diagnosis.
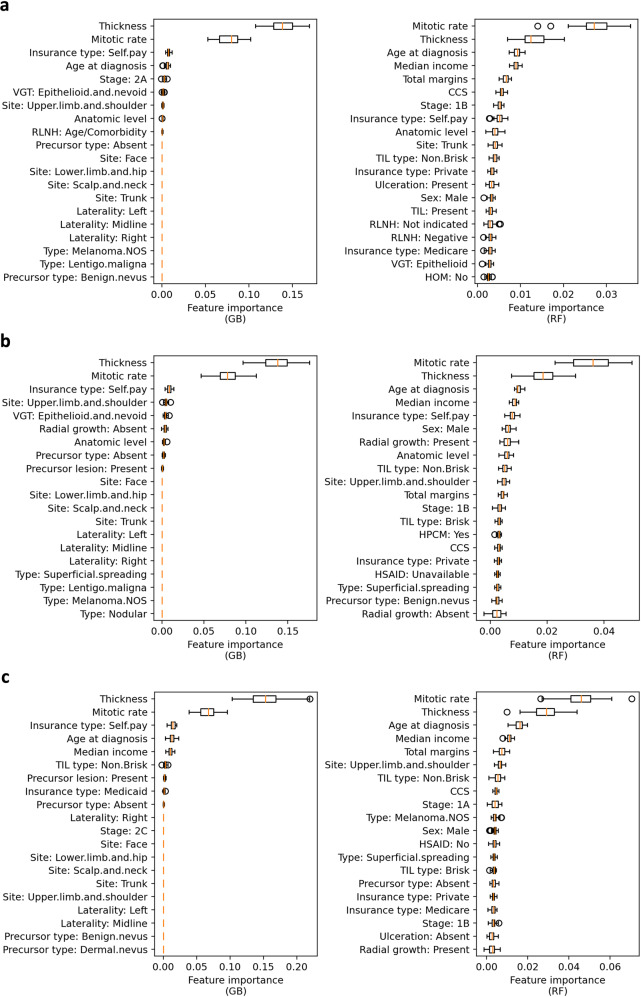
Permutation importance analysis across Gradient Boosting (GB) and Random Forest (RF) models consistently identified Breslow thickness and mitotic rate as the two dominant predictors — validating established pathologic risk factors while also surfacing novel signals such as TIL type and radial growth pattern.
The model also generates patient-level time-to-event recurrence trajectories, enabling individualized surveillance planning and counseling for patients with Stage I–II melanoma.
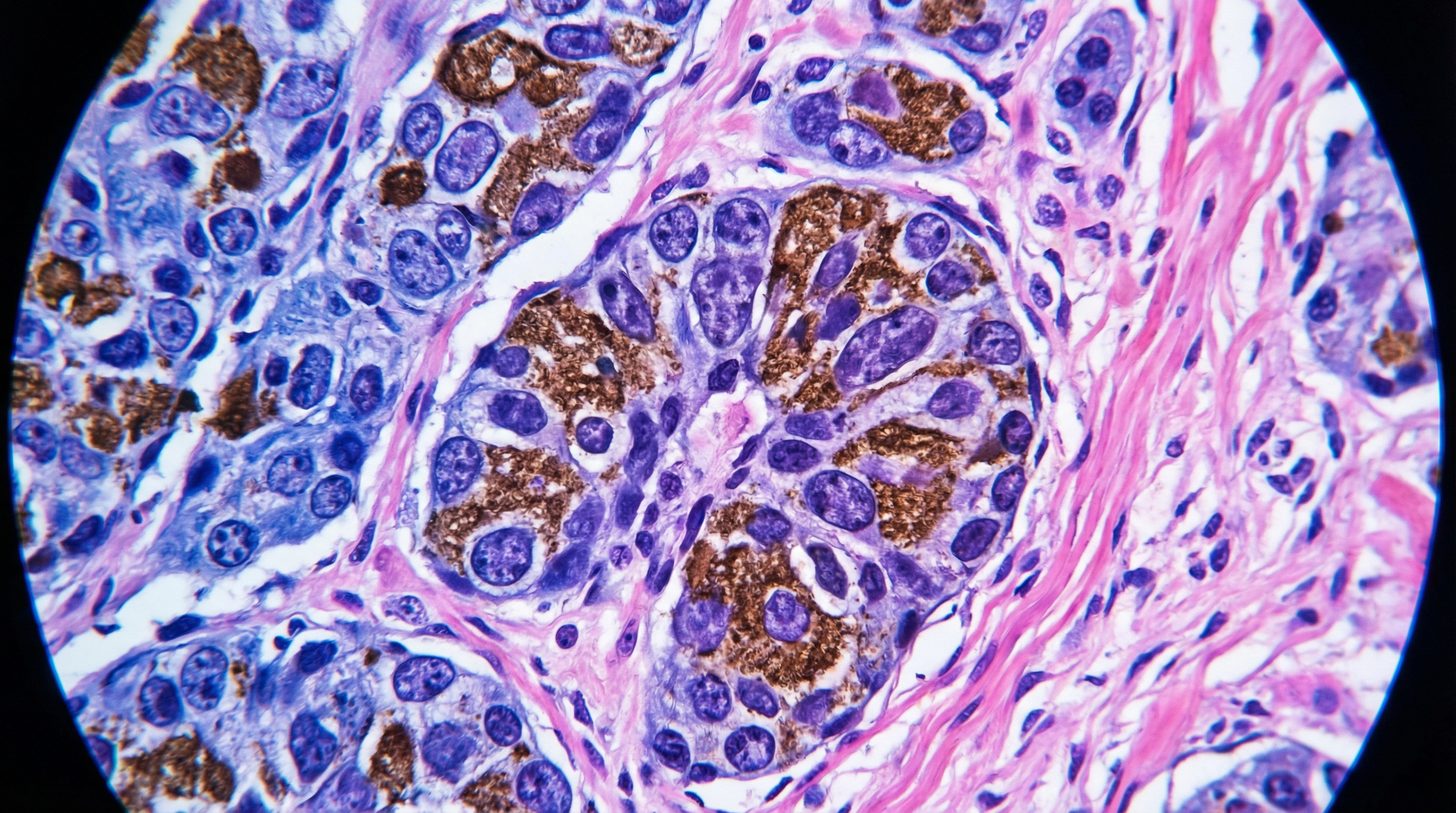
Our ongoing work focuses on integrating foundation models with attention-based multiple instance learning on whole-slide images to extract prognostic signal invisible to the human eye.
Our seminal population-level analysis of 8,637 ICI recipients and 8,637 matched controls established the true epidemiology of cutaneous immune-related adverse events (irAEs) — the spectrum of skin toxicities from checkpoint blockade and cancer immunotherapy, including rash, pruritus (itch), immune-mediated skin disease, inflammatory skin disease, vitiligo, and lichenoid eruptions — finding that only 10 of 43 previously reported dermatoses are actually elevated in ICI-treated patients.
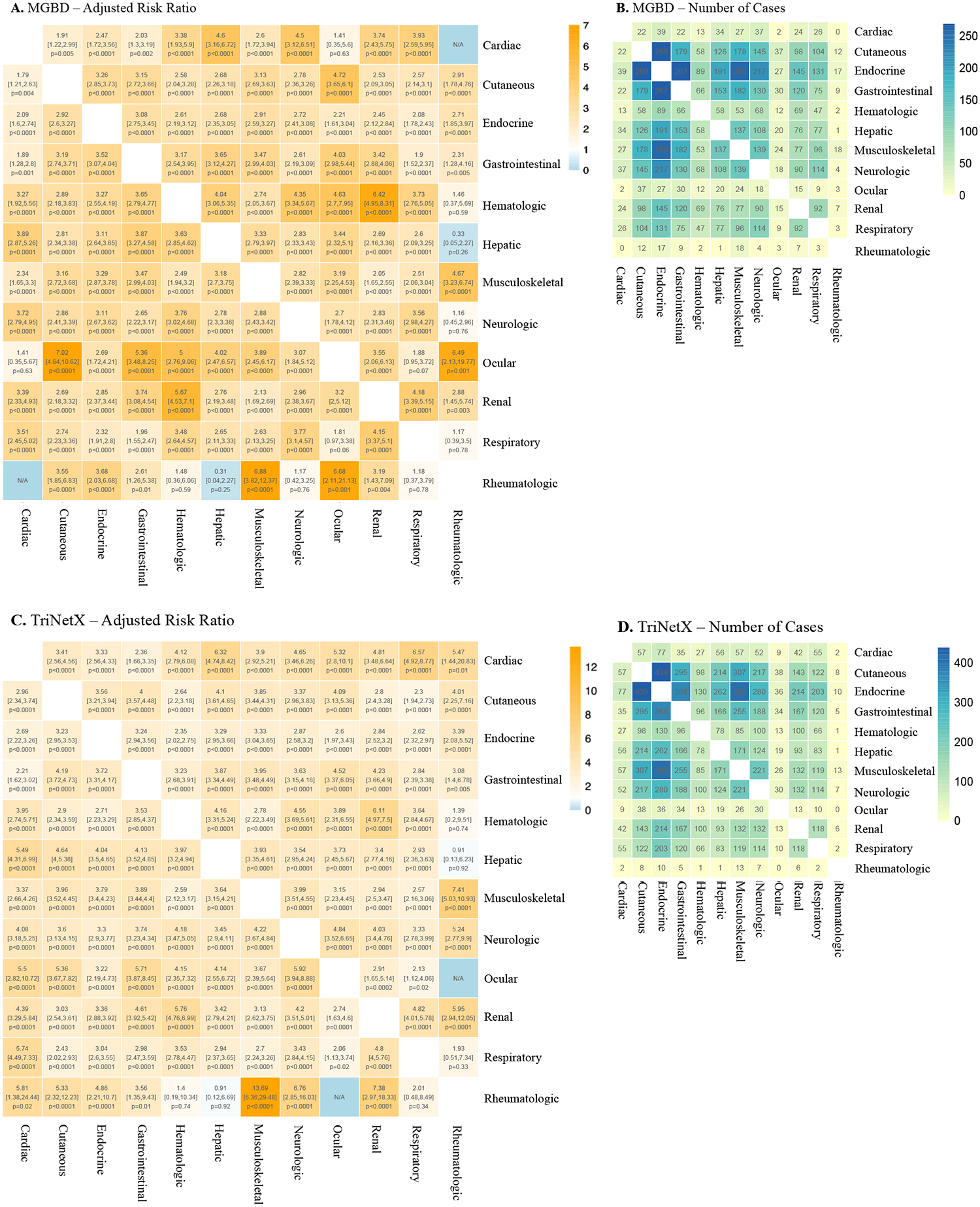
In our landmark Lancet Oncology 2024 study, we characterized the downstream implications of multi-organ irAEs across 12 organ systems in large MGB and TriNetX cohorts. Using non-negative matrix factorization, we identified seven distinct irAE clusters — finding that endocrine- and cutaneous-predominant clusters are associated with improved survival, while respiratory- and neurologic-predominant clusters confer worse outcomes.
Additional work across 14,016 patients demonstrated that cutaneous irAEs are associated with a 22% reduction in mortality, with specific morphologies (vitiligo, lichenoid, acneiform) predicting significantly improved survival — positioning irAEs as actionable biomarkers of immunotherapy response.
Our group has built one of the largest multi-institutional irAE registries and biorepositories, encompassing 24,000+ immunotherapy recipients treated between 2011 and 2025, leveraging AI in oncology and computational phenotyping to characterize toxicity patterns and treatment outcomes at scale. This registry also supports clinical trials in oncodermatology focused on immune-mediated skin disease.
To translate these findings into clinical practice, we co-led the development of consensus-based disease definitions for the diagnosis and severity grading of dermatologic irAEs (D-irAEs). Published in the Journal for ImmunoTherapy of Cancer (JITC) in 2024, this multi-institutional Delphi consensus — co-first authored by Chen and Semenov with a panel of 34 experts from 22 institutions — established standardized diagnostic criteria for the 10 most common D-irAE subtypes, enabling more precise classification across clinical and research settings.
Immunotherapy saves lives but its toxicities can be fatal — our registry and AI models are building the evidence base to predict, prevent, and personalize management of these complications.
In collaboration with the Broad Institute and Dana-Farber, we conducted the first large-scale GWAS of immune checkpoint inhibitor toxicity — identifying 3 genome-wide significant loci, including a variant near IL7 (p = 3.6×10⁻¹¹, HR 2.1) that colocalized with a cryptic exon affecting lymphocyte homeostasis.
Patients carrying this variant showed increased lymphocyte stability after ICI initiation, which was itself predictive of downstream irAEs and associated with improved survival — uncovering a mechanistic link between host autoimmunity and immunotherapy response.
Nature Medicine 2022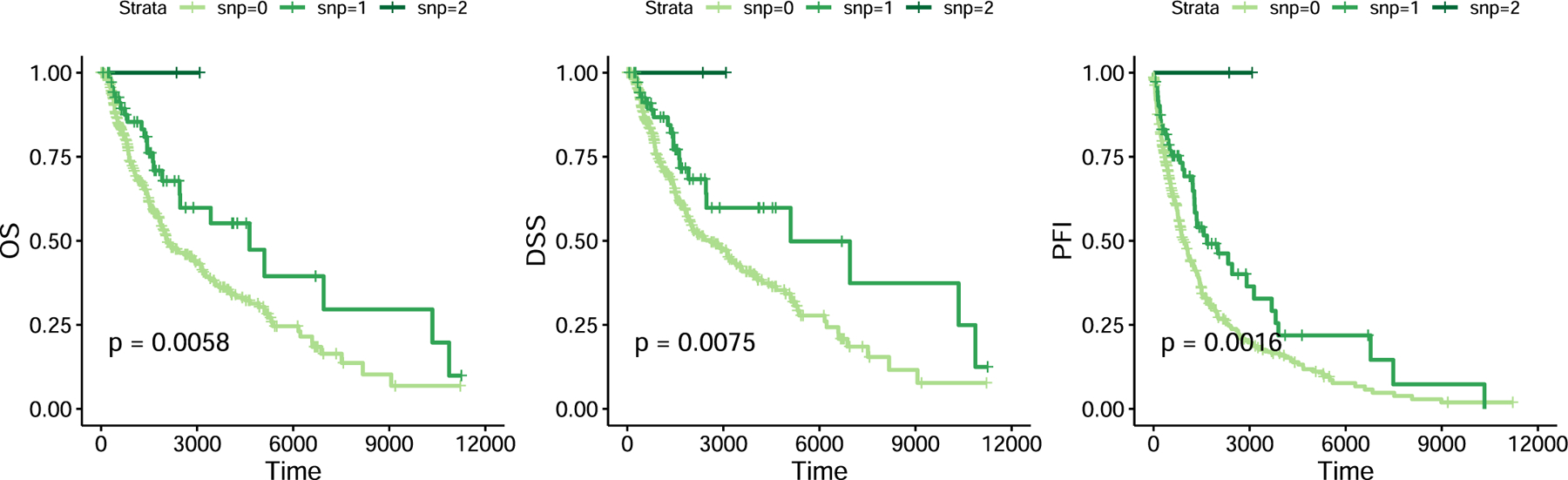
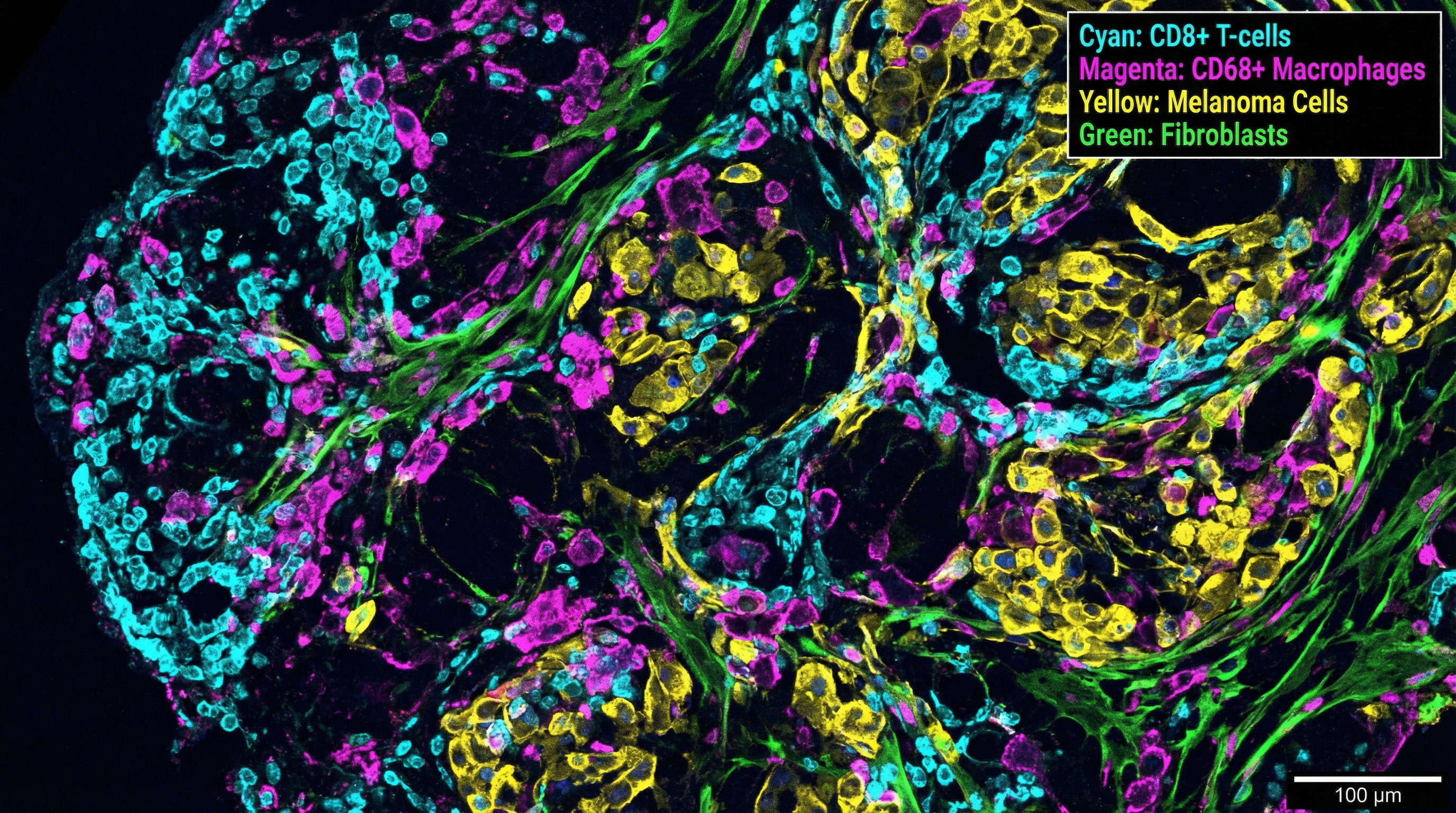
Our spatial biology program uses multiplex immunofluorescence imaging, spatial transcriptomics, and multiomic profiling of melanoma tissue to map cell-cell interactions within the tumor microenvironment — with a focus on in-transit melanoma as a model for studying stromal remodeling and immune evasion in cutaneous oncology.
By integrating single-cell and spatial resolution data, we identify biomarkers of treatment response, mechanisms of immune checkpoint blockade resistance, and spatially resolved signatures of melanoma progression — applying AI in skin disease at the tissue level.
We developed SpatialCells, an open-source software package for automated, region-based profiling of tumor microenvironments using spatially resolved multiplexed single-cell data — enabling high-throughput feature extraction and machine learning predictions from complex tissue specimens.
The tumor microenvironment determines whether immunotherapy succeeds or fails — our spatial tools reveal the cellular architecture that governs treatment response.